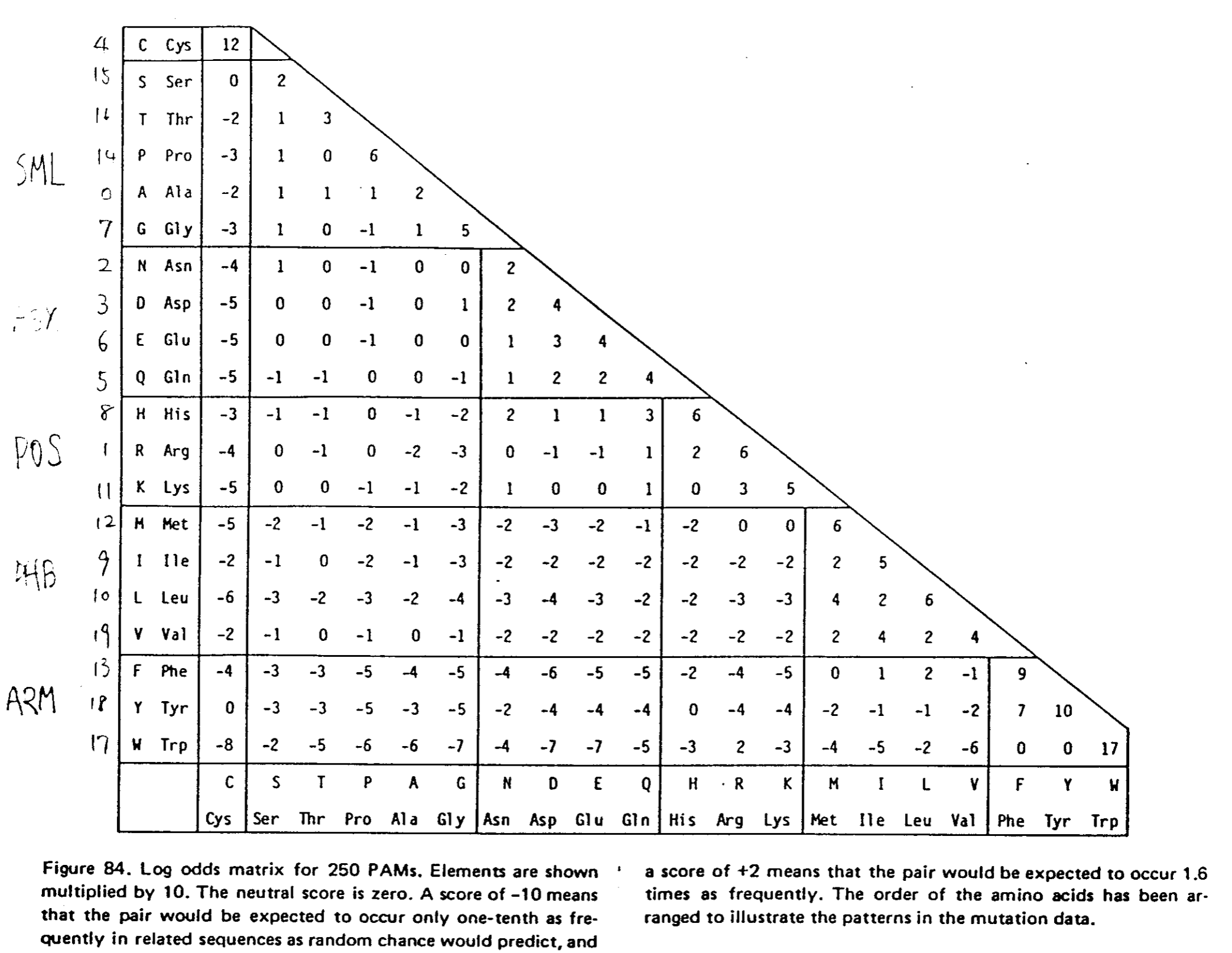
Matrices de scores de substitution : PAM¶
Rappel : Quelques diapositives sur l'alignement de 2 séquences et les matrices de substitution
Calcul d'une matrice de substitution PAM250 à partir d'alignements¶
Nous allons construire une matrice de type Dayhoff (la PAM250) pour l'alignement de séquences protéiques, ceci à partir d'alignements de séquences. La méthode utilise
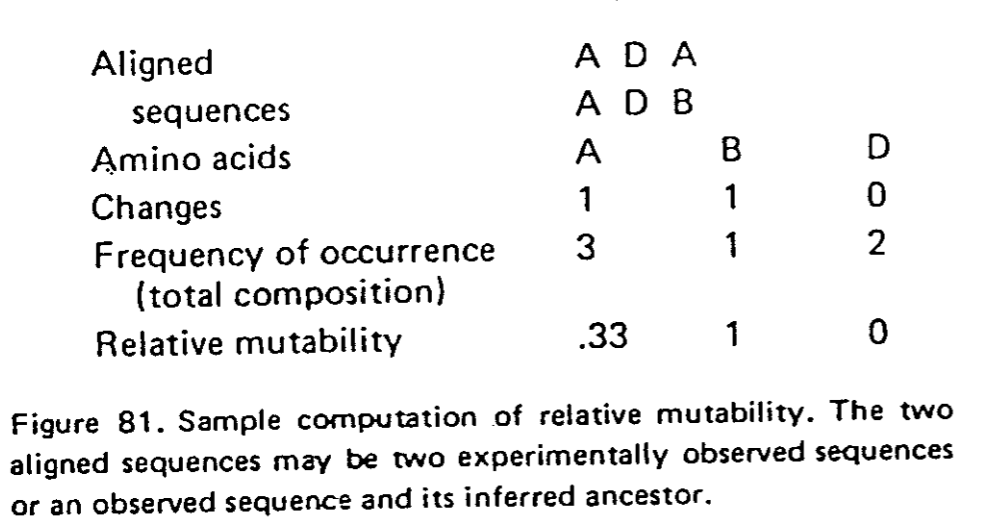
- une matrice comptage du nombre de substitutions observées pour chaque acide aminé (aa),
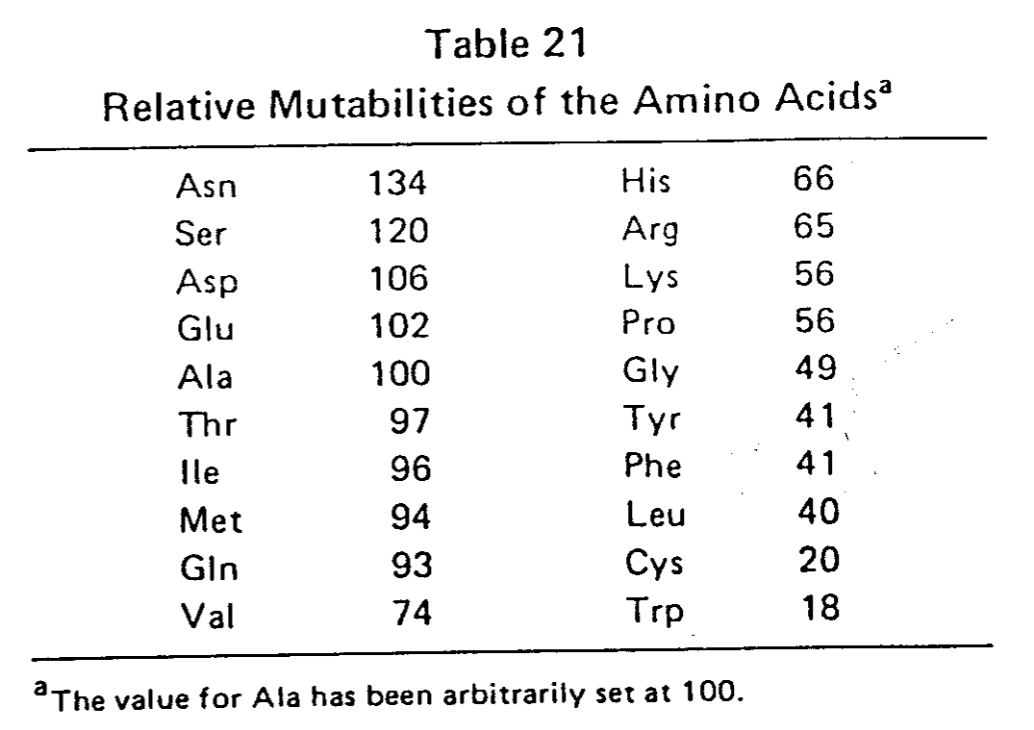
- la mutabilité de chaque aa,, c'est-à-dire la probabilité qu'un aa a de muter,
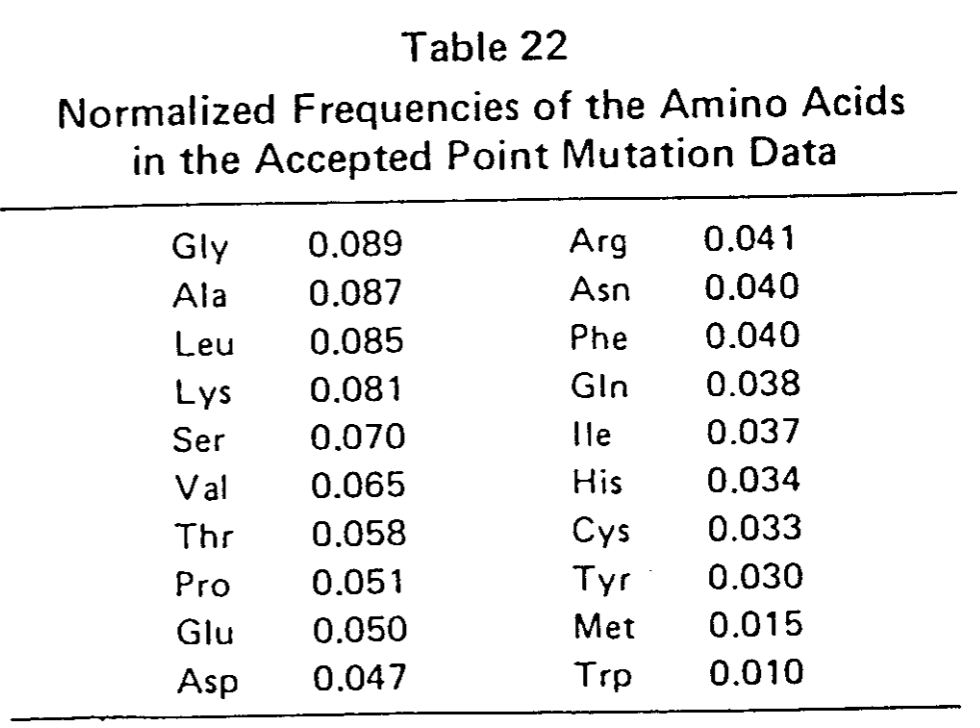
- la fréquence d'apparition de chaque aa dans une séquence quelconque
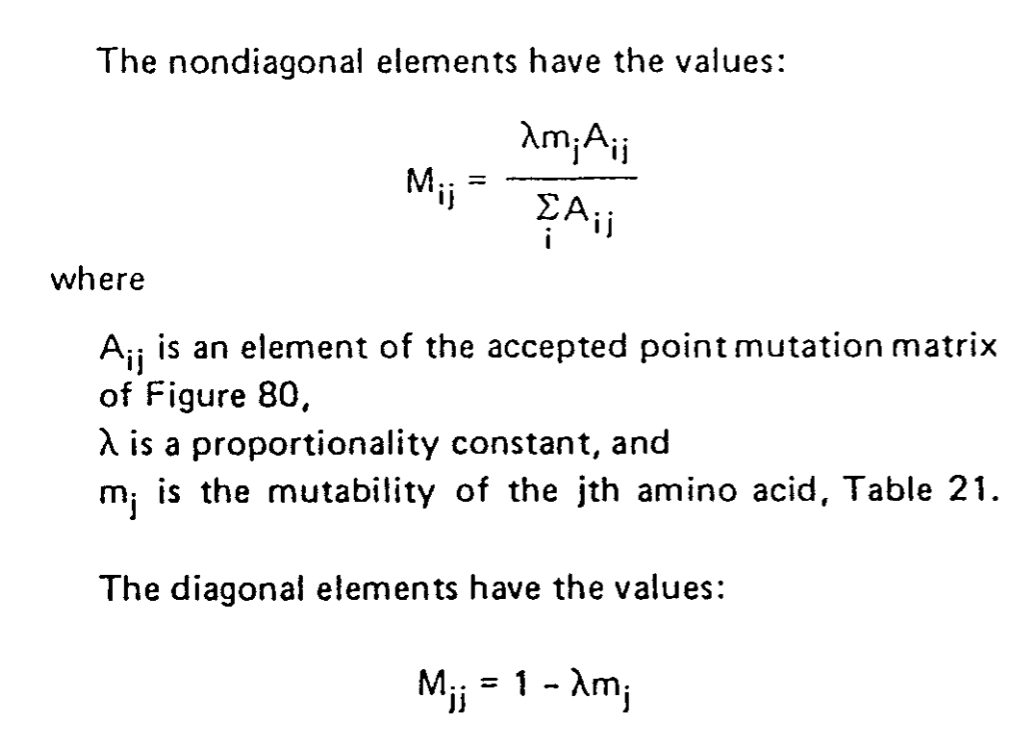
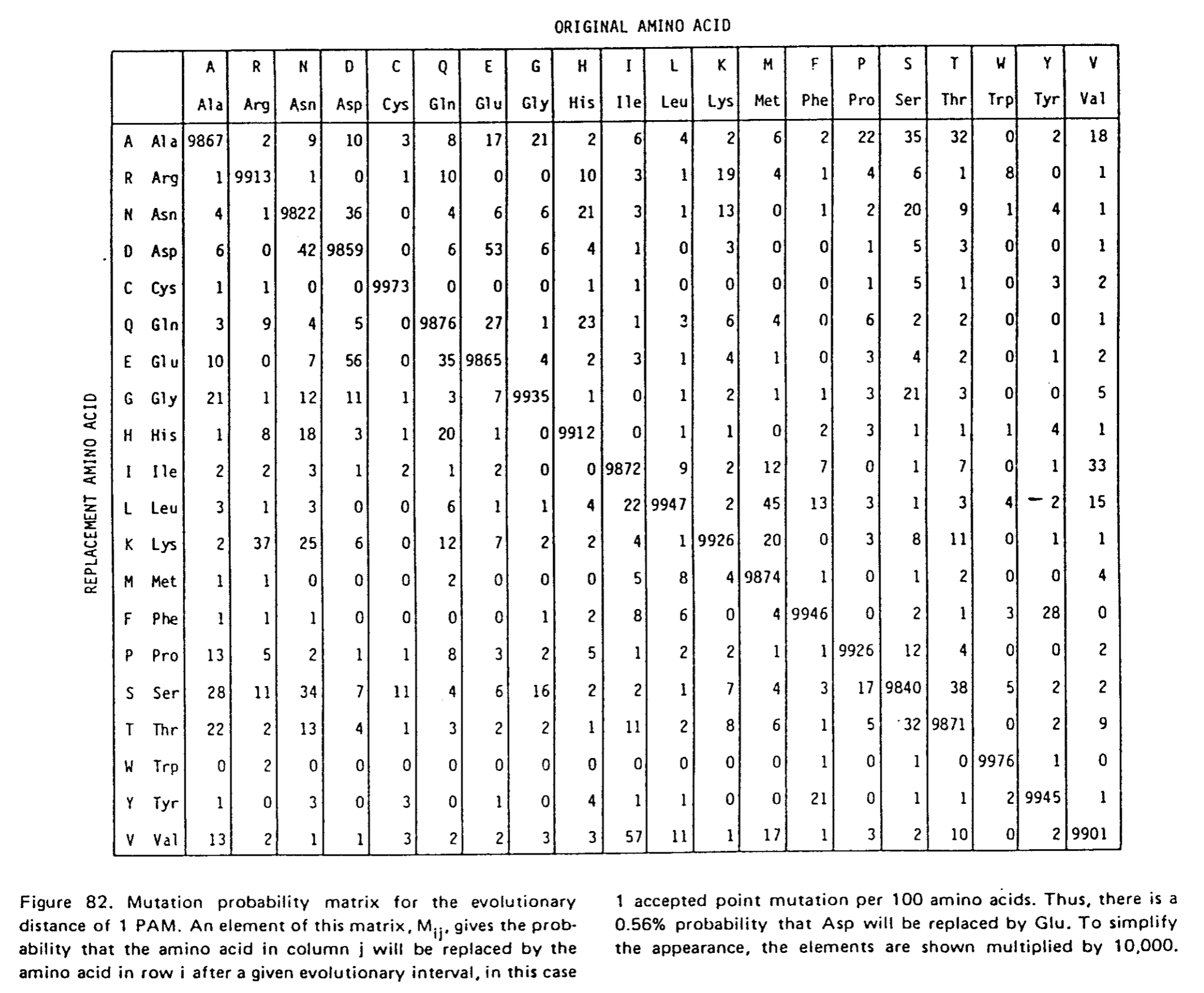
A partir de ces informations, la matrice de probabilité de mutations d'un aa en un autre peut-être calculée pour une distance évolutive correspondant au temps qu'il faut pour qu'une mutation sur 100 aa soit fixée dans la population.
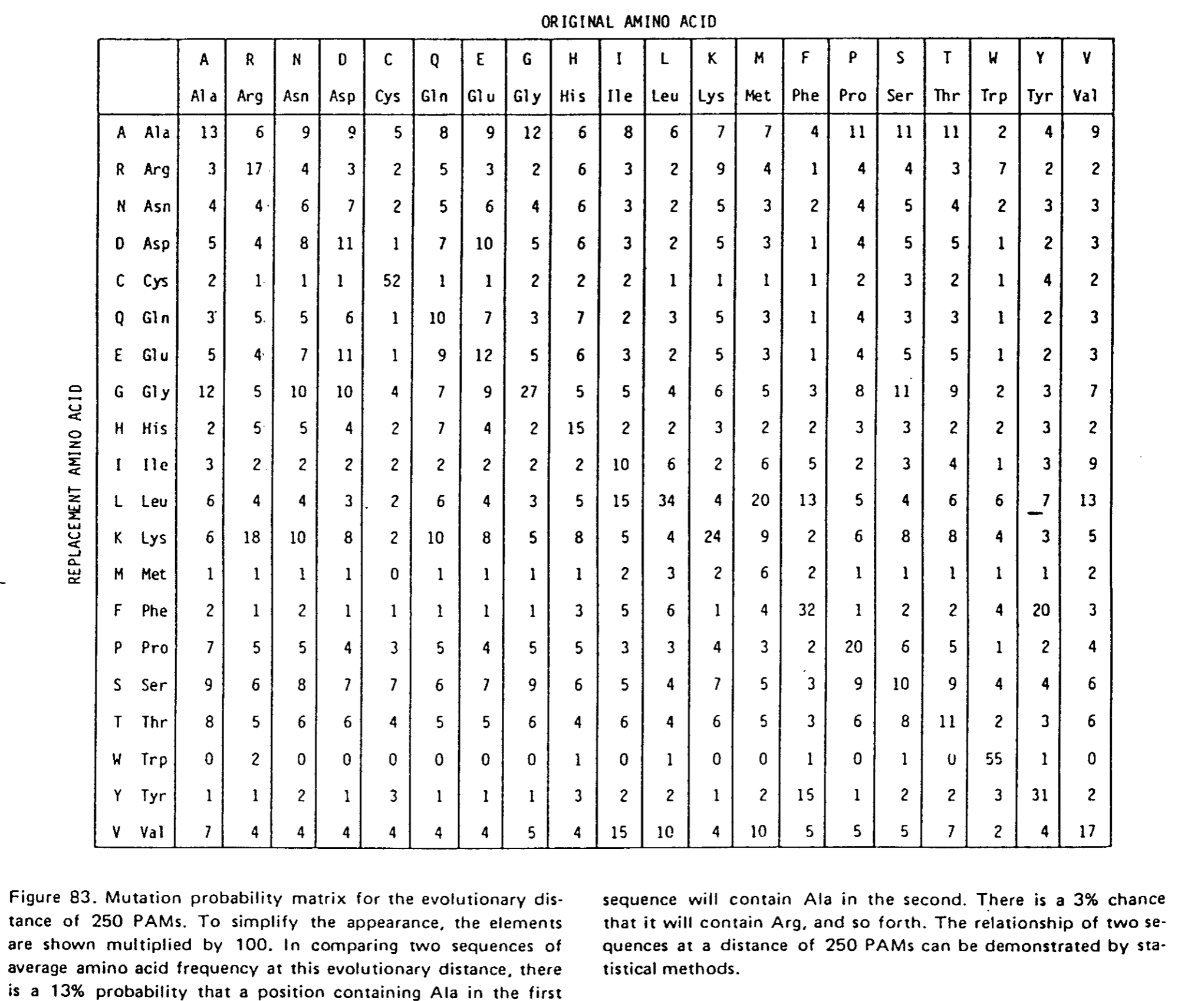
A partir de cette matrice, il est ensuite possible de calculer les matrices correspondant à des distances évolutives plus grandes. Et à partir de ces dernières matrices, on pourra calculer les matrices de scrores de substitutions utilisables pour faire l'alignement de séquences.
Nous allons donc suivre le cheminement décrit dans A Model of Evolutionary Change in Proteins, M.O Dayhoff, R.M. Schwartz and B.C. Orcutt, Atlas of Proteins Sequence and Structure, 1978
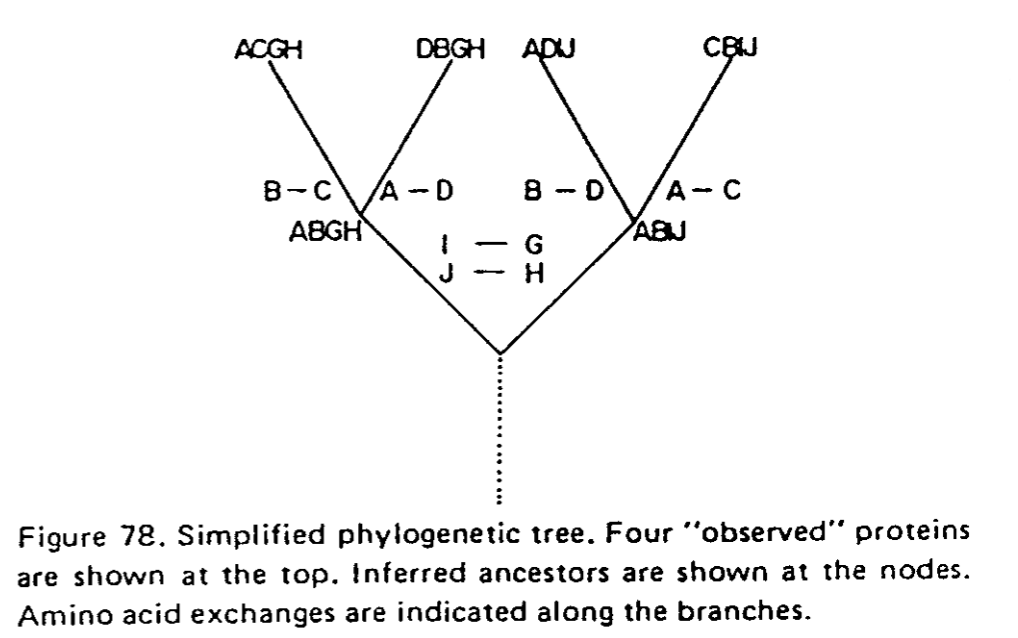
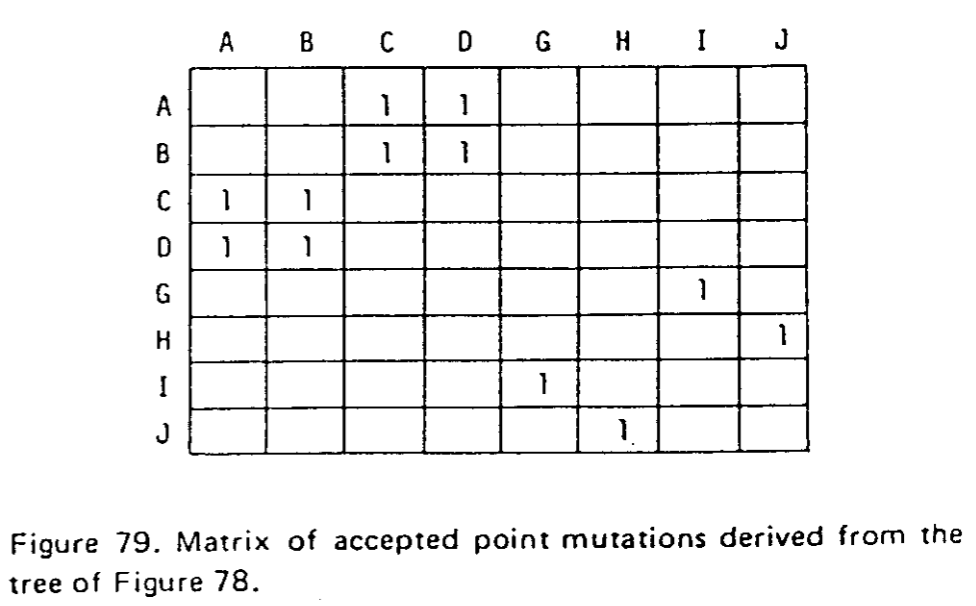
La première étape (déjà réalisée pour vous) consiste à réaliser les alignements globaux par familles de séquences, inférer les arbres de parenté correspondant afin de dénombrer le nombre de passage d'un acide aminé à un autre.
Matrice de comptage cm¶
Cette matrice de comptage correspond à la figure 80 :
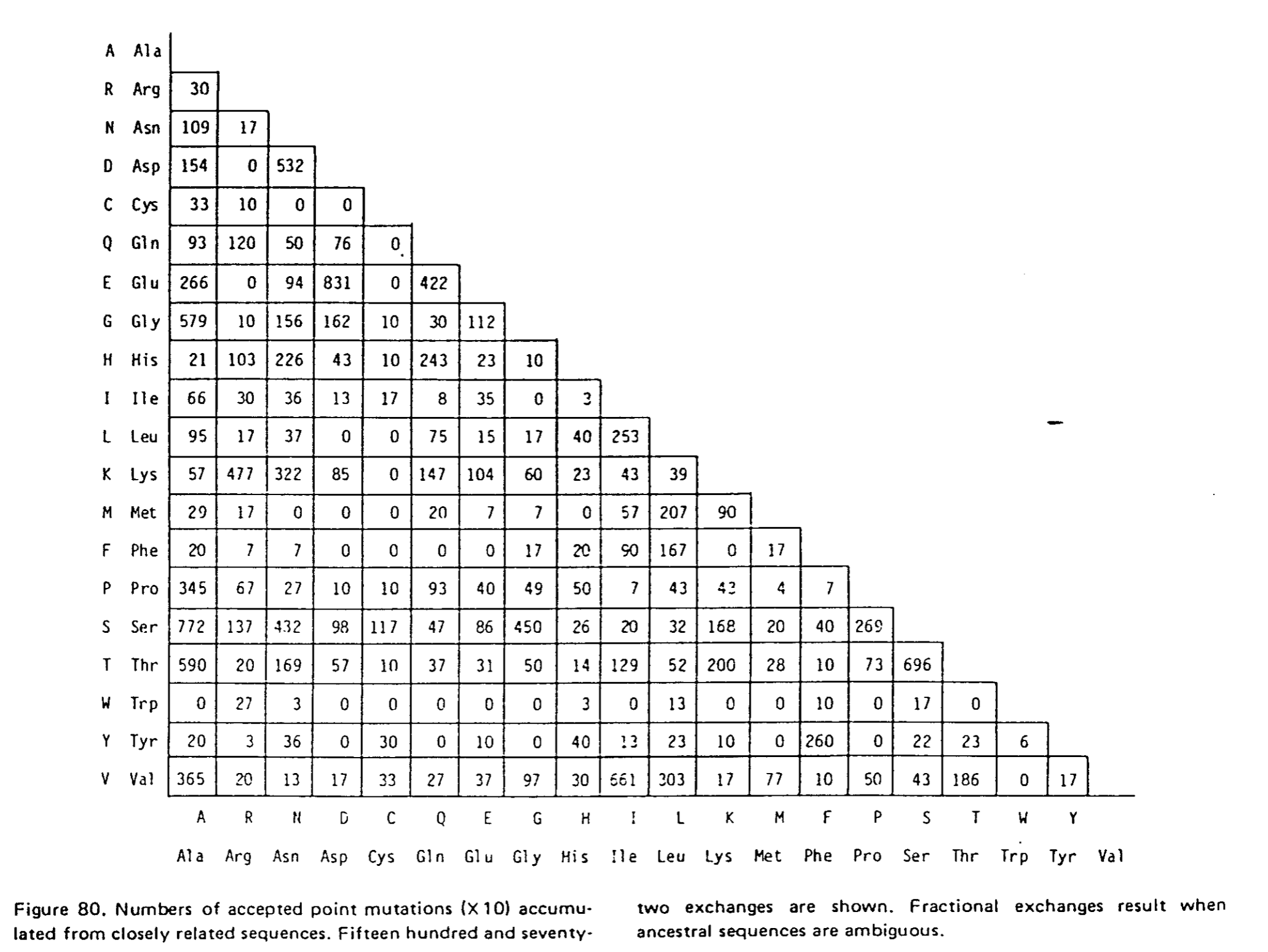
Cette matrice est symétrique.
Le fichier correspondant à la figure a été reconstitué : dayhoff.fig80.counts.csv.